This post expands on a previous post and gives more examples of harmonic morphisms to the tree . This graph is also called a star graph
on 3+1=4 vertices, or the bipartite graph
.

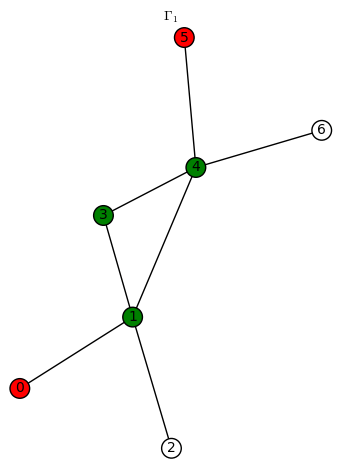
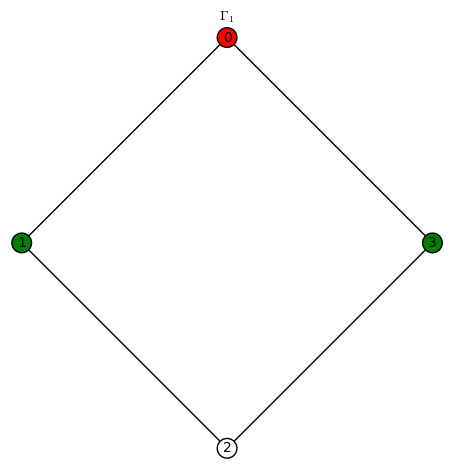
We indicate a harmonic morphism by a vertex coloring. An example of a harmonic morphism can be described in the plot below as follows: sends the red vertices in
to the red vertex of
(we let 3 be the numerical notation for the color red), the blue vertices in
to the blue vertex of
(we let 2 be the numerical notation for the color blue), the green vertices in
to the green vertex of
(we let 1 be the numerical notation for the color green), and the white vertices in
to the white vertex of
(we let 0 be the numerical notation for the color white).
First, a simple remark about harmonic morphisms in general: roughly speaking, they preserve adjacency. Suppose is a harmonic morphism. Let
be adjacent vertices of
. Then either (a)
and
“collapses” the edge (vertical)
or (b)
and the vertices
and
are adjacent in
. In the particular case of this post (ie, the case of
), this remark has the following consequence: since in
the green vertex is not adjacent to the blue or red vertex, none of the harmonic colored graphs below can have a green vertex adjacent to a blue or red vertex. In fact, any colored vertex can only be connected to a white vertex or a vertex of like color.
To get the following data, I wrote programs in Python using SageMath.
Example 1: There are only the 4 trivial harmonic morphisms , plus the “obvious” ones obtained from that below and those induced by permutations of the vertices:
 .
.
My guess is that the harmonic morphisms can be described in a similar manner. Likewise for the higher
graphs. Given a star graph
with a harmonic morphism to
, a leaf (connected to the center vertex 0) can be added to
and preserve “harmonicity” if its degree 1 vertex is colored 0. You can try to “collapse” such leafs, without ruining the harmonicity property.
Example 2: For graphs like

there are only the 4 trivial harmonic morphisms , plus the “obvious” ones obtained from that above and those induced by permutations of the vertices with a non-zero color.
Example 2.5: Likewise, for graphs like

there are only the 4 trivial harmonic morphisms , plus the “obvious” ones obtained from that above and those induced by permutations of the vertices with a non-zero color.
Example 3: This is really a non-example. There are no harmonic morphisms from the (3-dimensional) cube graph (whose vertices are those of the unit cube) to .
More generally, take two copies of a cyclic graph on n vertices, , one hovering over the other. Now, connect each vertex of the top copy to the corresponding vertex of the bottom copy. This is a cubic graph that can be visualized as a “thick” regular polygon. (The cube graph is the case
.) I conjecture that there is no harmonic morphism from such a graph to
.
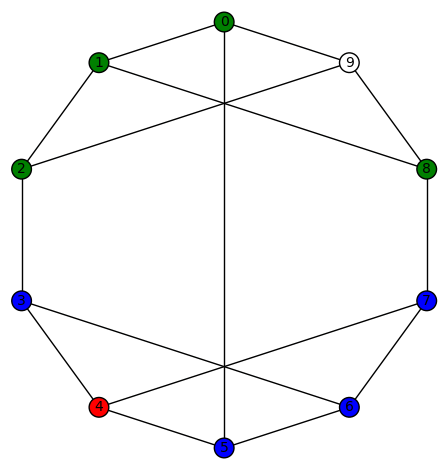
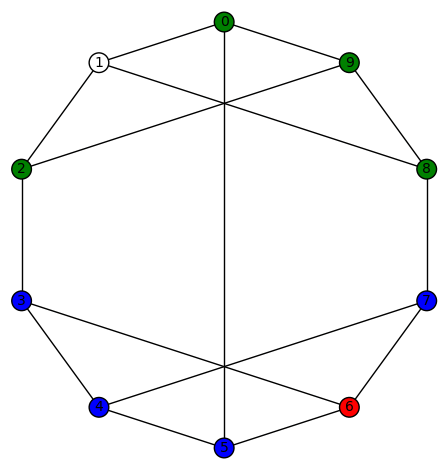
Example 4: There are 30 non-trivial harmonic morphisms for the Peterson graph (the last of the 19 simple cubic graphs on 10 vertices listed on this wikipedia page). Here is an example:

Another interesting fact is that this graph has an automorphism group (isomorphic to the symmetric group on 5 letters) which acts transitively on the vertices.
Example 5: There are 12 non-trivial harmonic morphisms for the complete bipartite (“utility”) graph
. They are all obtained from either

or

by permutations of the vertices with a non-zero color (3!+3!=12).
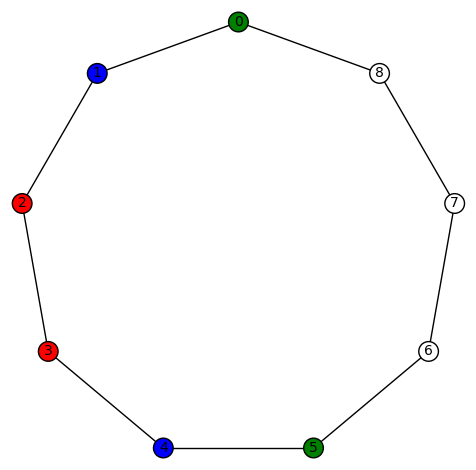
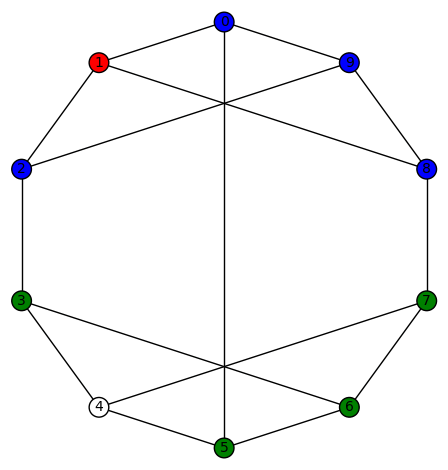
Example 6: There are 6 non-trivial harmonic morphisms for the cubic graph
, where
and
. This graph has diameter 3, girth 3, and edge-connectivity 3. It’s automorphism group is size 4, generated by (5,9) and (1,8)(2,7)(3,6). The harmonic morphisms are all obtained from
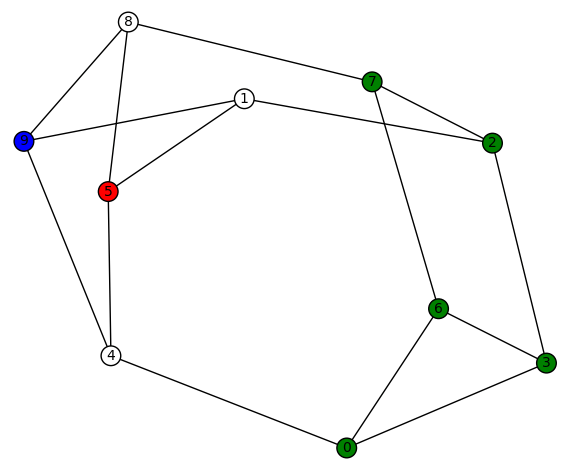
by permutations of the vertices with a non-zero color (3!=6). This graph might be hard to visualize but it is isomorphic to the simple cubic graph having LCF notation [−4, 3, 3, 5, −3, −3, 4, 2, 5, −2]:

which has a nice picture. This is the ninth of the 19 simple cubic graphs on 10 vertices listed on this wikipedia page.
Example 7: (a) The first of the 19 simple cubic graphs on 10 vertices listed on this wikipedia page is the graph depicted as:
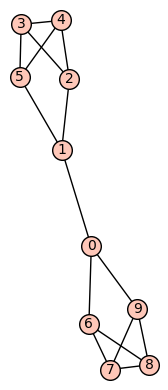
This graph has diameter 5, automorphism group generated by (7,8), (6,9), (3,4), (2,5), (0,1)(2,6)(3,7)(4,8)(5,9). There are no non-trivial harmonic morphisms .
(b) The second of the 19 simple cubic graphs on 10 vertices listed on this wikipedia page is the graph depicted as:

This graph has diameter 4, girth 3, automorphism group generated by (7,8), (0,5)(1,2)(6,9). There are no non-trivial harmonic morphisms .
(c) The third of the 19 simple cubic graphs on 10 vertices listed on this wikipedia page is the graph depicted as:

This graph has diameter 3, girth 3, automorphism group generated by (4,5), (0,1)(8,9), (0,8)(1,9)(2,7)(3,6). There are no non-trivial harmonic morphisms .
Example 8: The fourth of the 19 simple cubic graphs on 10 vertices listed on this wikipedia page is the graph depicted as:

This graph has diameter 3, girth 3, automorphism group generated by (4,6), (3,5), (1,8)(2,7)(3,4)(5,6), (0,9). There are 12 non-trivial harmonic morphisms . For example,

and the remaining (3!=6 total) colorings obtained by permutating the non-zero colors. Another example is
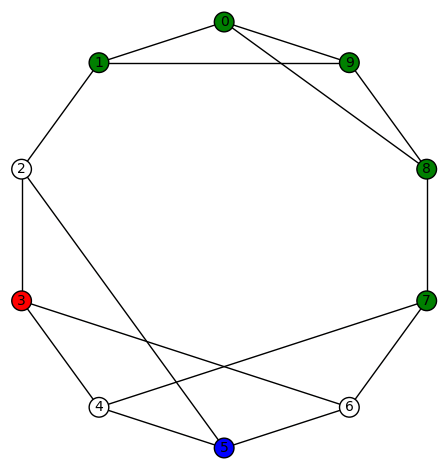
and the remaining (3!=6 total) colorings obtained by permutating the non-zero colors.
Example 9: (a) The fifth of the 19 simple cubic graphs on 10 vertices listed on this wikipedia page is the graph depicted as:
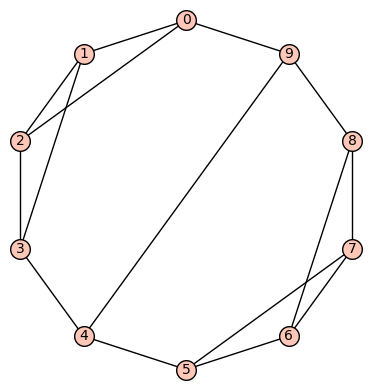
Its SageMath command is Gamma1 = graphs.LCFGraph(10,[2,2,-2,-2,5],2) There are no non-trivial harmonic morphisms .
(b) The sixth of the 19 simple cubic graphs on 10 vertices listed on this wikipedia page is the graph depicted as:
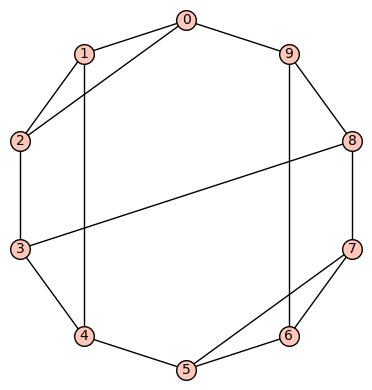
Its SageMath command is Gamma1 = graphs.LCFGraph(10,[2,3,-2,5,-3],2) There are no non-trivial harmonic morphisms .
Example 10: The seventh of the 19 simple cubic graphs on 10 vertices listed on this wikipedia page is the graph depicted as:

Its SageMath command is Gamma1 = graphs.LCFGraph(10,[2,3,-2,5,-3],2). Its automorphism group is order 12, generated by (1,2)(3,7)(4,6), (0,1)(5,6)(7,9), (0,4)(1,6)(2,5)(3,9). There are 6 non-trivial harmonic morphisms , each obtained from the one above by permuting the non-zero colors.
Example 11: The eighth of the 19 simple cubic graphs on 10 vertices listed on this wikipedia page is the graph depicted as:

Its SageMath command is Gamma1 = graphs.LCFGraph(10,[5, 3, 5, -4, -3, 5, 2, 5, -2, 4],1). Its automorphism group is order 2, generated by (0,3)(1,4)(2,5)(6,7). There are no non-trivial harmonic morphisms .
Example 12: (a) The tenth (recall the 9th was mentioned above) of the 19 simple cubic graphs on 10 vertices listed on this wikipedia page is the graph depicted as:
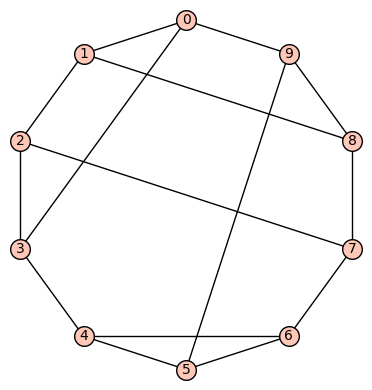
Its SageMath command is Gamma1 = graphs.LCFGraph(10,[3, -3, 5, -3, 2, 4, -2, 5, 3, -4],1). Its automorphism group is order 6, generated by (2,8)(3,9)(4,5), (0,2)(5,6)(7,9). There are no non-trivial harmonic morphisms .
(b) The 11th of the 19 simple cubic graphs on 10 vertices listed on this wikipedia page is the graph depicted as:

Its SageMath command is Gamma1 = graphs.LCFGraph(10,[-4, 4, 2, 5, -2],2). Its automorphism group is order 4, generated by (0,1)(2,9)(3,8)(4,7)(5,6), (0,5)(1,6)(2,7)(3,8)(4,9). There are no non-trivial harmonic morphisms .
(c) The 12th of the 19 simple cubic graphs on 10 vertices listed on this wikipedia page is the graph depicted as:
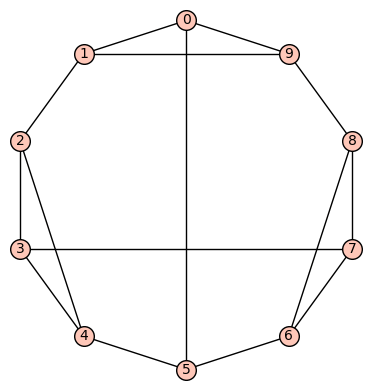
Its SageMath command is Gamma1 = graphs.LCFGraph(10,[5, -2, 2, 4, -2, 5, 2, -4, -2, 2],1). Its automorphism group is order 6, generated by (1,9)(2,8)(3,7)(4,6), (0,4,6)(1,3,8)(2,7,9). There are no non-trivial harmonic morphisms .
(d) The 13th of the 19 simple cubic graphs on 10 vertices listed on this wikipedia page is the graph depicted as:
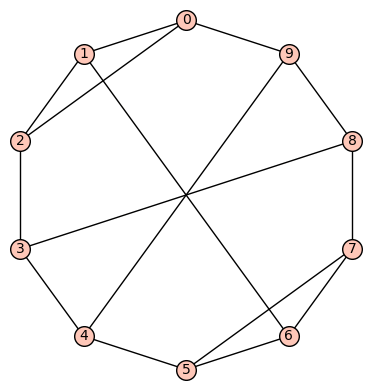
Its SageMath command is Gamma1 = graphs.LCFGraph(10,[2, 5, -2, 5, 5],2). Its automorphism group is order 8, generated by (4,8)(5,7), (0,2)(3,9), (0,5)(1,6)(2,7)(3,8)(4,9). There are no non-trivial harmonic morphisms .
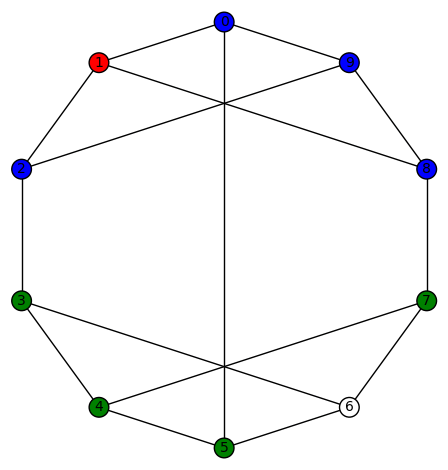
Example 13: The 14th of the 19 simple cubic graphs on 10 vertices listed on this wikipedia page is the graph depicted as:

By permuting the non-zero colors, we obtain 3!=6 harmonic morphisms from this one. Another harmonic morphism is depicted as:

By permuting the non-zero colors, we obtain 3!=6 harmonic morphisms from this one. And another harmonic morphism is depicted as:

By permuting the non-zero colors, we obtain 3!=6 harmonic morphisms from this one. Its SageMath command is Gamma1 = graphs.LCFGraph(10,[5, -3, -3, 3, 3],2). Its automorphism group is order 48, generated by (4,6), (2,8)(3,7), (1,9), (0,2)(3,5), (0,3)(1,4)(2,5)(6,9)(7,8). There are a total of 18=3!+3!+3! non-trivial harmonic morphisms .
Example 14: The 15th of the 19 simple cubic graphs on 10 vertices listed on this wikipedia page is the graph depicted as:

By permuting the non-zero colors, we obtain 3!=6 harmonic morphisms from this one. Its SageMath command is Gamma1 = graphs.LCFGraph(10,[5, -4, 4, -4, 4],2). Its automorphism group is order 8, generated by (2,7)(3,8), (1,9)(2,3)(4,6)(7,8), (0,5)(1,4)(2,3)(6,9)(7,8). There are a total of 6=3! non-trivial harmonic morphisms .
Example 15: (a) The 16th of the 19 simple cubic graphs on 10 vertices listed on this wikipedia page is the graph depicted as:

Its SageMath command is Gamma1 = graphs.LCFGraph(10,[5, -4, 4, 5, 5],2). Its automorphism group is order 4, generated by (0,3)(1,2)(4,9)(5,8)(6,7), (0,5)(1,6)(2,7)(3,8)(4,9). There are no non-trivial harmonic morphisms .
(b) The 17th of the 19 simple cubic graphs on 10 vertices listed on this wikipedia page is the graph depicted as:

Its SageMath command is Gamma1 = graphs.LCFGraph(10,[5, 5, -3, 5, 3],2). Its automorphism group is order 20, generated by (2,6)(3,7)(4,8)(5,9), (0,1)(2,5)(3,4)(6,9)(7,8), (0,2)(1,9)(3,5)(6,8). This group acts transitively on the vertices. There are no non-trivial harmonic morphisms .
(c) The 18th of the 19 simple cubic graphs on 10 vertices listed on this wikipedia page is the graph depicted as:
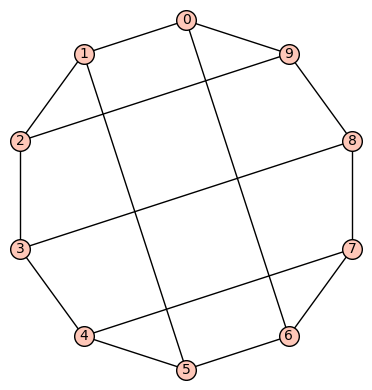
This is an example of a “thick polygon” graph, already mentioned in Example 3 above. Its SageMath command is Gamma1 = graphs.LCFGraph(10,[-4, 4, -3, 5, 3],2). Its automorphism group is order 20, generated by (2,5)(3,4)(6,9)(7,8), (0,1)(2,6)(3,7)(4,8)(5,9), (0,2)(1,9)(3,6)(4,7)(5,8). This group acts transitively on the vertices. There are no non-trivial harmonic morphisms .
(d) The 19th in the list of 19 is the Petersen graph, already in Example 4 above.
We now consider some examples of the cubic graphs having 12 vertices. According to the House of Graphs there are 109 of these, but we use the list on this wikipedia page.
Example 16: I wrote a SageMath program that looked for harmonic morphisms on a case-by-case basis. If there is no harmonic morphism then, instead of showing a graph, I’ll list the edges (of course, the vertices are 0,1,…,11) and the SageMath command for it.
, where
.
SageMath command:
V1 = [0,1,2,3,4,5,6,7,8,9,10,11]
E1 = [(0, 1), (0, 2), (0, 11), (1, 2), (1, 6), (2, 3), (3, 4), (3, 5), (4, 5), (4, 6), (5, 6), (7, 8), (7, 9), (7, 11), (8, 9), (8, 10), (9, 10), (10, 11)]
Gamma1 = Graph([V1,E1])
(Not in LCF notation since it doesn’t have a Hamiltonian cycle.)
, where
.
SageMath command:
V1 = [0,1,2,3,4,5,6,7,8,9,10,11]
E1 = [(0, 1), (0, 6), (0, 11), (1, 2), (1, 3), (2, 3), (2, 5), (3, 4), (4, 5), (4, 6), (5, 6), (7, 8), (7, 9), (7, 11), (8, 9), (8, 10), (9, 10), (10, 11)]
Gamma1 = Graph([V1,E1])
(Not in LCF notation since it doesn’t have a Hamiltonian cycle.)
, where
.
SageMath command:
V1 = [0,1,2,3,4,5,6,7,8,9,10,11]
E1 = [(0,1),(0,3),(0,11),(1,2),(1,6),(2,3),(2,5),(3,4),(4,5),(4,6),(5,6),(7,8),(7,9),(7,11),(8,9),(8,10),(9,10),(10,11)]
Gamma1 = Graph([V1,E1])
(Not in LCF notation since it doesn’t have a Hamiltonian cycle.)
- This example has 12 non-trivial harmonic morphisms.
SageMath command:
V1 = [0,1,2,3,4,5,6,7,8,9,10,11]
E1 = [(0,1),(0,3),(0,11),(1,2),(1,6),(2,3),(2,5),(3,4),(4,5),(4,6),(5,6),(7,8),(7,9),(7,11),(8,9),(8,10),(9,10),(10,11)]
Gamma1 = Graph([V1,E1])
(Not in LCF notation since it doesn’t have a Hamiltonian cycle.) We show two such morphisms:

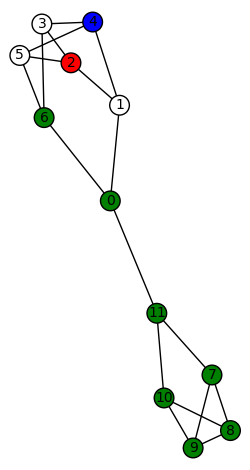
The other non-trivial harmonic morphisms are obtained by permuting the non-zero colors. There are 3!=6 for each graph above, so the total number of harmonic morphisms (including the trivial ones) is 6+6+4=16.
, where
.
SageMath command:
Gamma1 = graphs.LCFGraph(12, [3, -2, -4, -3, 4, 2], 2)
- This example has 12 non-trivial harmonic morphisms.
, where
. (This only differs by one edge from the one above.)
SageMath command:
Gamma1 = graphs.LCFGraph(12, [3, -2, -4, -3, 3, 3, 3, -3, -3, -3, 4, 2], 1)
We show two such morphisms:

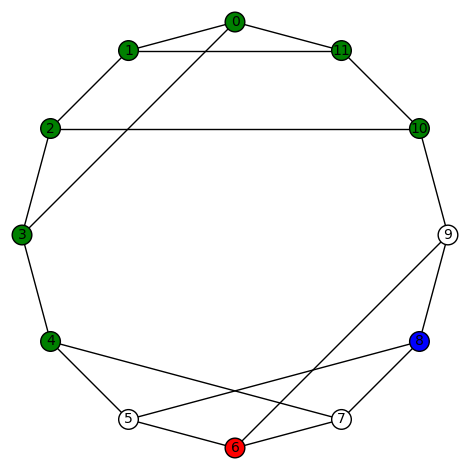
And here is another plot of the last colored graph:
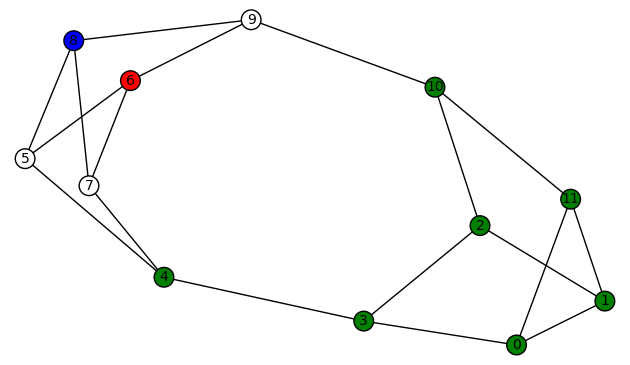
The other non-trivial harmonic morphisms are obtained by permuting the non-zero colors. There are 3!=6 for each graph above, so the total number of harmonic morphisms (including the trivial ones) is 6+6+4=16.
, where
.
SageMath command:
Gamma1 = graphs.LCFGraph(12, [4, 2, 3, -2, -4, -3, 2, 3, -2, 2, -3, -2], 1)
- This example has 48 non-trivial harmonic morphisms.
, where
.
SageMath command:
Gamma1 = graphs.LCFGraph(12, [3, 3, 3, -3, -3, -3], 2)
This example is also interesting as it has a large number of automorphisms – its automorphism group is size 64, generated by (8,10), (7,9), (2,4), (1,3), (0,5)(1,2)(3,4)(6,11)(7,8)(9,10), (0,6)(1,7)(2,8)(3,9)(4,10)(5,11). Here are examples of some of the harmonic morphisms using vertex-colored graphs:


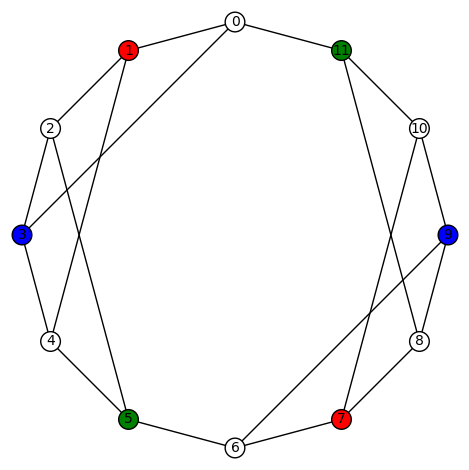

I think all the other non-trivial harmonic morphisms are obtained by (a) permuting the non-zero colors, or (b) applying a element of the automorphism group of the graph.
- (list under construction)




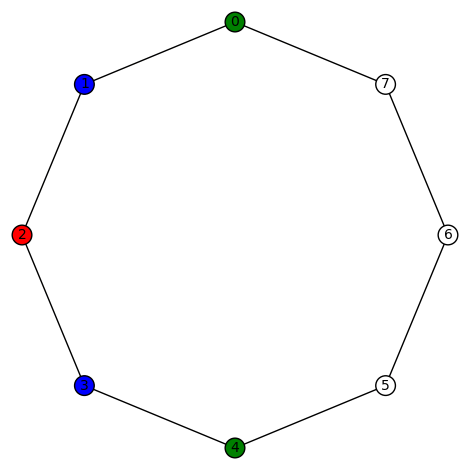
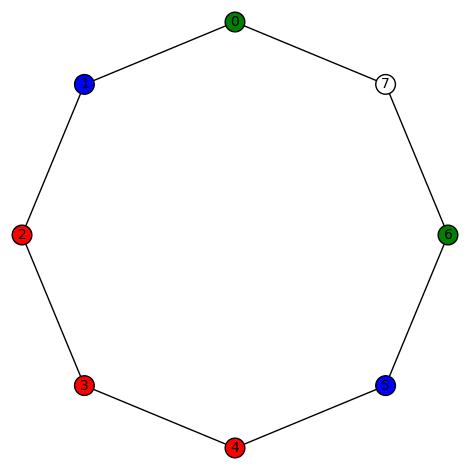
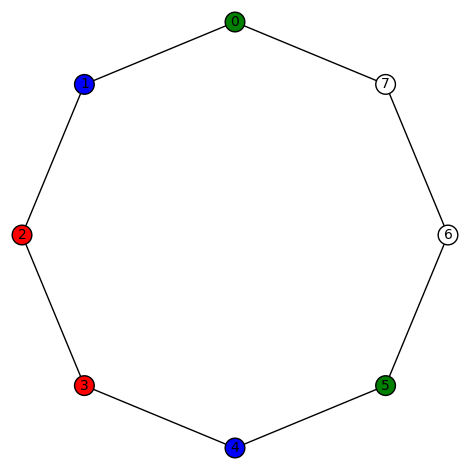











You must be logged in to post a comment.